 
Yet, this effect cannot be rationalized and it is often specific to the particular optimized tRNA. Multiple rounds of random nucleotide mutagenesis in the tRNA body 16 and combinatorial changes of different tRNA segments 17 have been shown to improve suppression efficiency, implying that other tRNA elements, in addition to the anticodon, can modulate this efficiency. However, through the exchange of the anticodon triplet only few natural tRNAs can be repurposed into suppressor tRNAs 10, 11, 12, 13, 14, 15, generating tRNAs with fairly modest effectivity in decoding stop codons and correcting nonsense mutations. In a few species, the detrimental effects of pervasive nonsense mutations are kept low by suppressor tRNAs, which commonly arise by mutation in a tRNA’s anticodon 9 to decode the newly arising stop codon. Nonsense mutations within protein-coding sequences convert a sense triplet into a stop codon, which in humans is connected to various devastating pathologies 7, 8. At a mechanistic level, the nucleophilic reactions in the ribosomal peptidyl-transferase center in the case of sense-codon recognition by an aminoacyl-tRNA and RF-mediated hydrolysis of peptidyl-tRNA are markedly different 6. Two decoding RFs are present in bacteria, RF1 and RF2, whereas all three stop codons are recognized by one factor, eRF1/aRF1 in eukaryotes and archaea, respectively 5. However, three codons, UAA, UAG, and UGA, are almost exclusively reserved for terminating translation across the three kingdoms of life 4, and are instead, decoded by proteins termed release factors (RFs). 
In nature, the 61 triplet codons encoding the 20 amino acids in proteins are decoded by, on average, 40–46 distinct transfer RNAs (tRNAs or isoacceptors) in bacteria, 41–55 cytosolic and 22 mitochondrial tRNAs in eukaryotes 1, 2, 3. The systematic analysis, combined with structural insights, provides a rationale for targeted repurposing of tRNAs to correct devastating nonsense mutations that introduce a premature stop codon. In the context of the suppressor tRNA, the conformation of the UGA codon resembles that of a sense-codon rather than when canonical translation termination release factors are bound, suggesting conformational flexibility of the stop codons dependent on the nature of the A-site ligand. We determine the structure of the ribosome in a complex with the designed tRNA bound to a UGA stop codon in the A site at 2.9 Å resolution. Optimization within the TΨC-stem to stabilize binding to the elongation factor, displays the most potent effect in enhancing suppression activity. 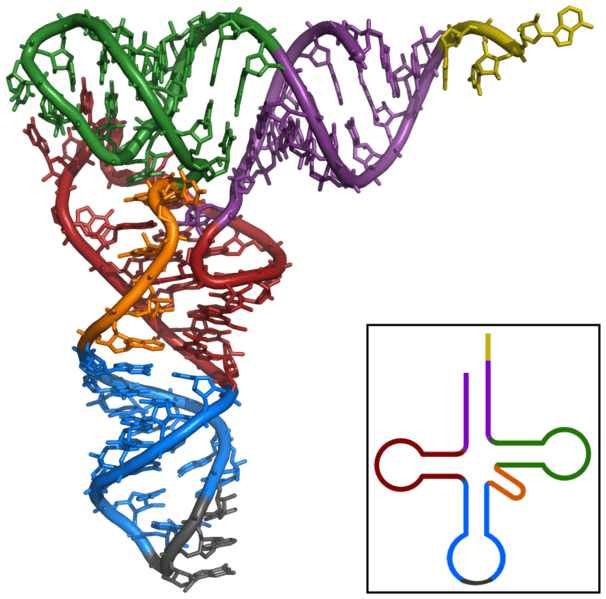
The tRNA designs harness various functionally conserved aspects of sense-codon decoding tRNAs. Here, we design de novo transfer RNAs (tRNAs) that efficiently decode UGA stop codons in Escherichia coli. Three stop codons (UAA, UAG and UGA) terminate protein synthesis and are almost exclusively recognized by release factors. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
